  #Python vectorize for loop code+ 1 and for i in 1:3 y = x + 1 end compile down to almost exactly the same code and perform comparably. 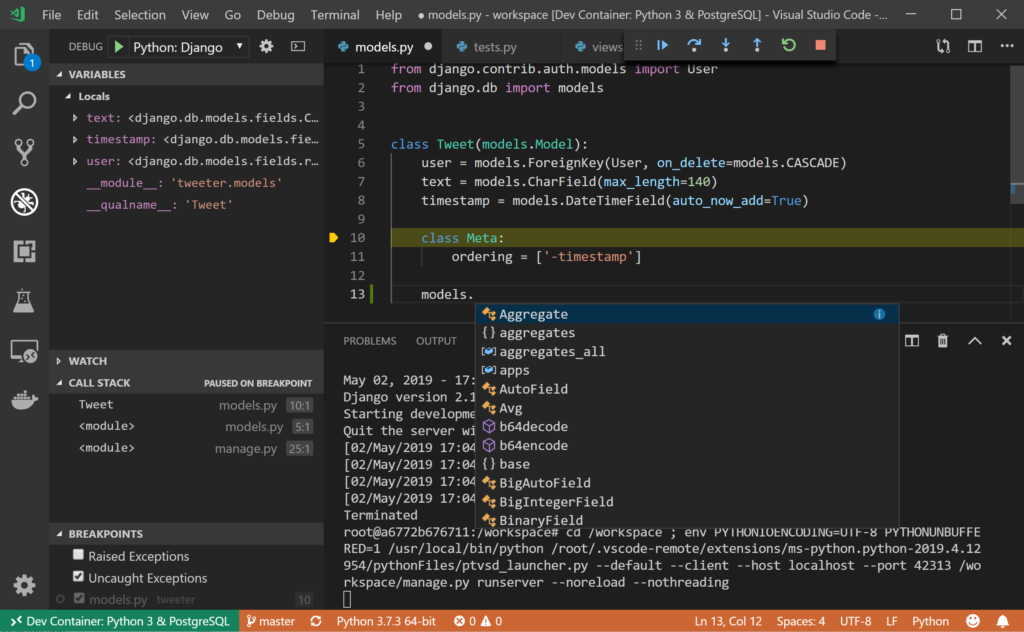
So in Python and Matlab, it’s faster to “vectorize” your code by, for example, only paying the cost of looking up the + operation once for the entire vector x rather than once for each element x. has an inherent overhead from the way the language works. Every loop iteration, every call to +, every array lookup, etc. This kind of vectorization is helpful in languages like Python and Matlab because every operation in Python is slow. When we talk about “vectorized” code in Python/Numpy/Matlab/etc., we are usually referring to the fact that code like: x = There are two meanings of the word “vectorization” in common usage, and they refer to different things. I think there is some PR in llvm underway that teaches min/max commutativity and associativity to llvm. The above is much faster and uglier than what we had before. Source: Long discussions on github when the above code got merged. The above code is a compromise that is still correct, and reliably uses 128 bit wide simd and maybe sometimes gets 256 bit simd right, but is unfortunately quite ugly and probably suboptimal on weak arm. And third, due to the way is implemented, there is currently no sensible way of opting out of julia-correct handling of corner cases, in order to get the blazingly fast pure simd minimum/ maximum. It can optimize integer arithmetic and float products and sums, but fails at float min/max reassociation (regardless of Secondly, the x86 min/max instructions on floating points have subtly different behavior with respect to NaN (and maybe signed zeros?) than julia. Unfortunately, llvm is notoriously bad at figuring this out. They all subtly mismatch!įor example, julia’s min/ max (as well as naive mathematical min/max) are associative and commutative, which is the best possible world for clever reorderings. 
In between, there is the compiler (llvm), the floating point semantics offered by the cpu, and the floating point semantics guaranteed by julia. For some operations, there are even built-in reductions running on a (128-512 bit) vector. For many operations, the cpu can operate on large vectors (128-512 bit) at the same time, and this is especially useful if these vectors are contiguous in memory. “sequential execution” is a lie: Your CPU is actually a very complex, badly documented and highly parallel beast that interprets the high-level language known as “assembly”. Hence, a*(b*(c*d)) is often slower than (a*b)*(c*d) (for generalized reducers like *, +, min, max, hash, etc). The simple answer (even before simd) boils down to dependency chains: Your CPU can do a lot of work in every single cycle, but most operations take multiple cycles until their result is ready. How does using four variables instead of one allow simd optimizations to take place? The variable names suggest that it has something to do with simd. V4 = v4 || return for i in start:4:simdstop Julia by default compiles down to machine code, why then is this implementation of reduce operation faster than the naive implementation: function mapreduce_impl(f, op::Union, 
In case of numpy, vectorization offloads heavy computation to the code in compiled libraries, improving speed. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
